App Note I May 17, 2017
List of protein hydrodynamic diameters DH
(i) Experimental protein diameters are analyzed from switching dynamics measurements with standard 48 bp DNA nanolevers (length = 16 nm) using the drift-diffusion ‘Lollipop’ model described in J. Phys. Chem. B 118, 597 (2014).
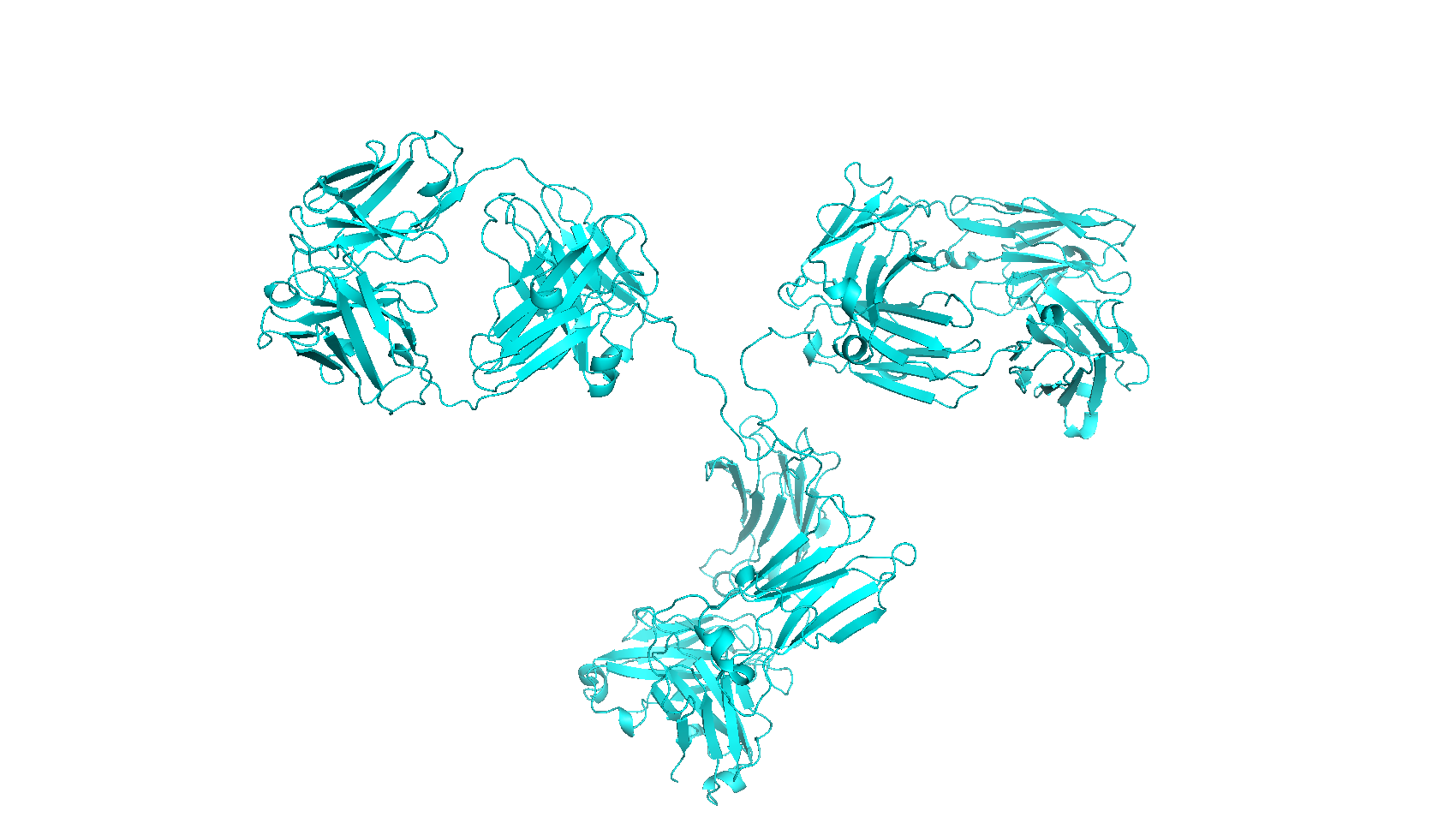
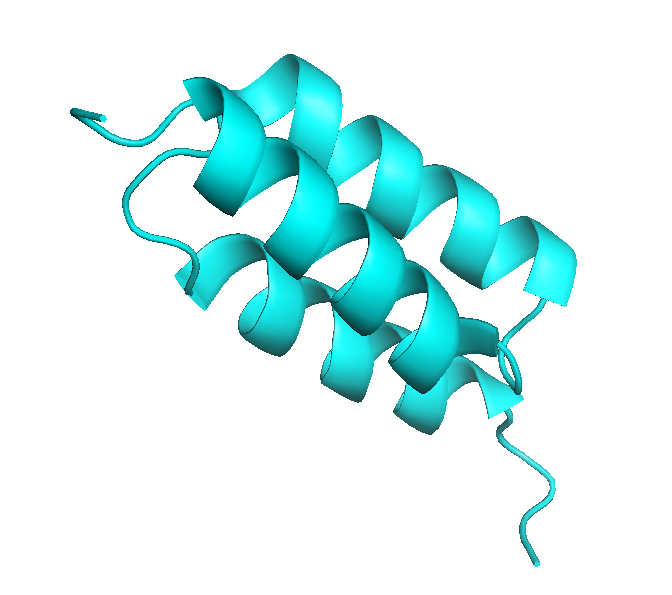
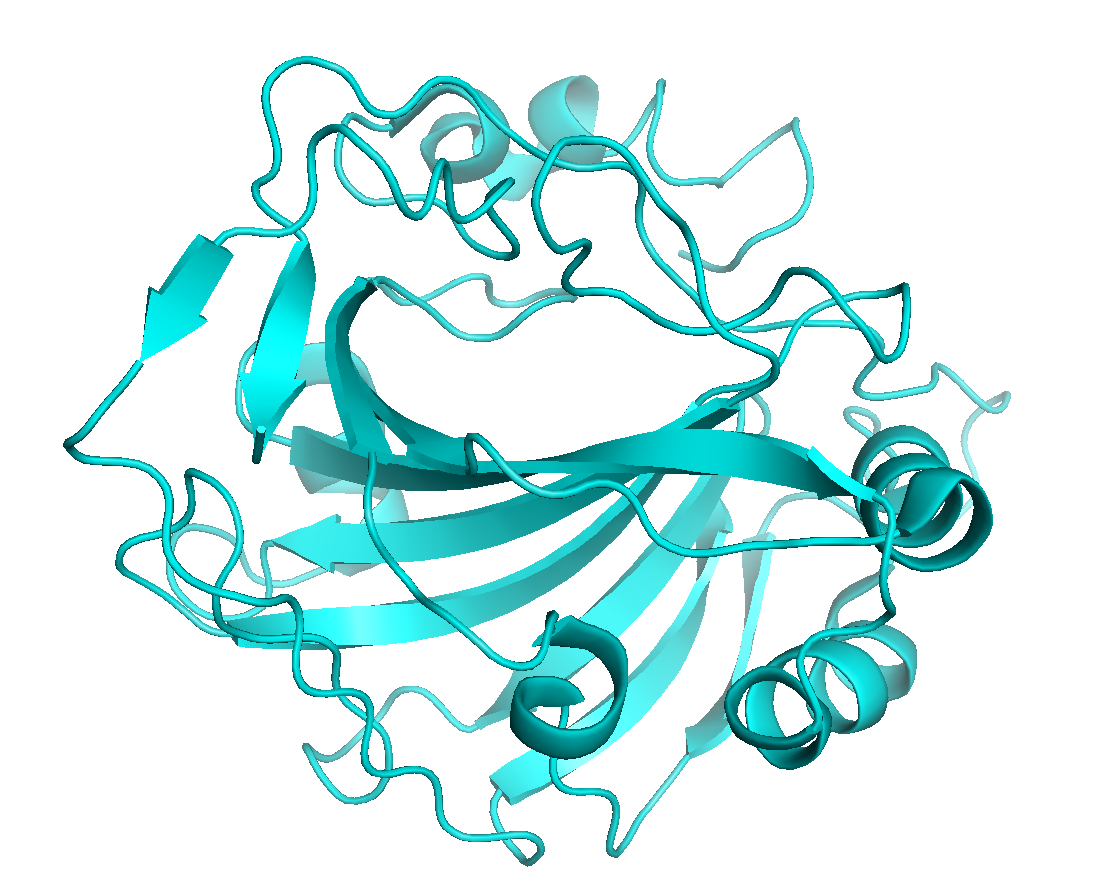
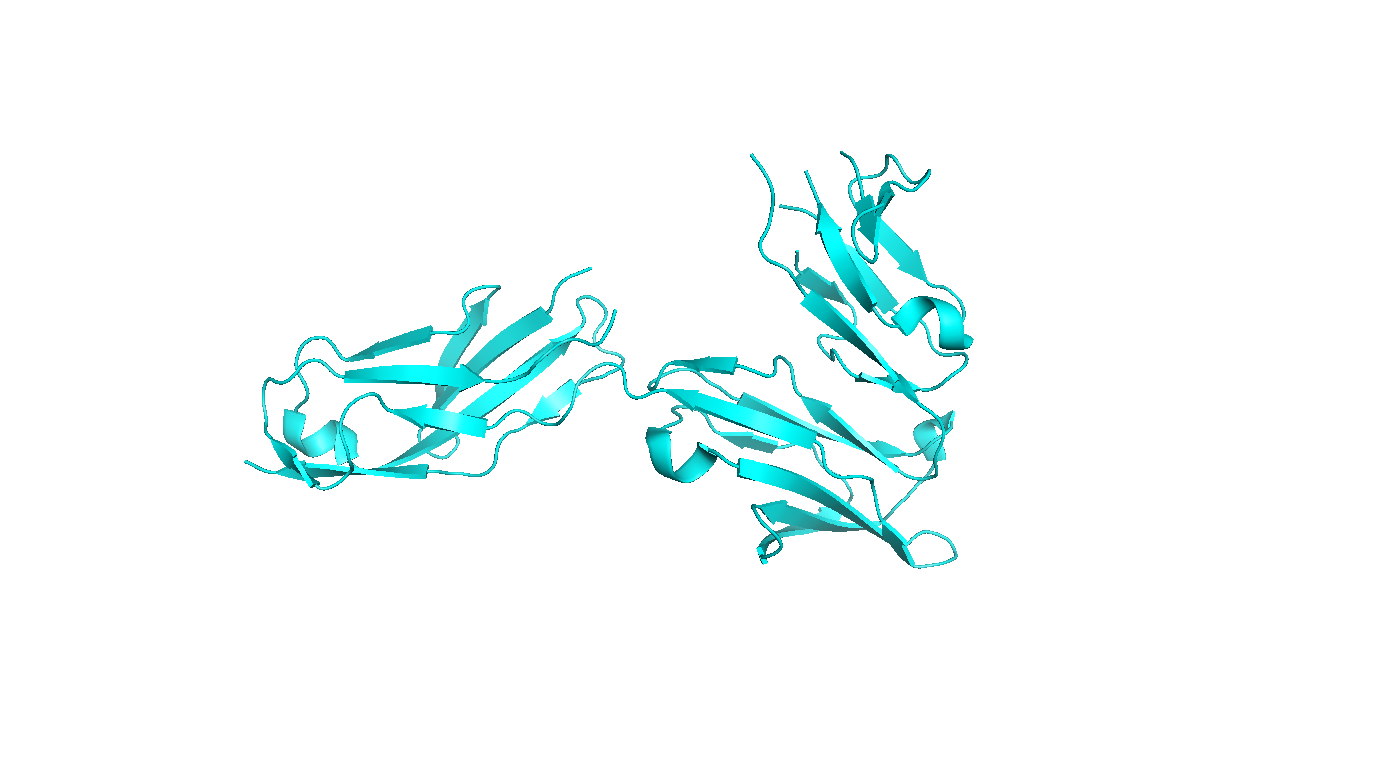
(ii) Theoretical calculations of protein diameters from PDB files are performed with the HydroPro algorithm [Garcia de la Torre et al. Biophys. J. 78:179, (2000)], that calculates an effective hydrodynamic diameter from atomic coordinates using a hydrodynamic bead-shell model. Note that crystal structures may not always reflect the native protein structure in solution (especially for large proteins with several domains connected by flexible linkers).