switchSENSE®
eLearning Center
We cover an array of various, different applications.
Browse through publications, application notes,
tutorials, and more.
Molecule Type
Analytical Method

Biophysical Analysis of CRBN Variants in PROTAC Ternary Kinetics

DISCOVER bispecific antibodies: generation, biophysical characterization and functional validation

scIC Binding kinetics on cells

DISCOVER reverse transcriptase activity: catalytic rates of nucleic acid-modifying enzymes

DISCOVER small molecule-induced protein conformational changes using DNA origami nanolevers

Insight into the avidity–affinity relationship of the bivalent, pH-dependent interaction between IgG and FcRn

DISCOVER screening of PROTACs/molecular glues: ternary complex analysis with induced proximity

Real-time measurement of DNA polymerase activity and inhibition with switchSENSE® and the heliX® biosensor

Reverse transcription as key step in RNA in vitro evolution with unnatural base pairs

Repurposing the mammalian RNA-binding protein Musashi-1 as an allosteric translation repressor in bacteria

Balancing the Affinity and Tumor Cell Binding of a Two-in-One Antibody Simultaneously Targeting EGFR and PD-L1

DISCOVER therapeutic antibodies: kinetics of antibody-drug conjugates measured directly on cells

DISCOVER protein-DNA conjugate purification: Effortless conjugate preparation, purification, and analysis with the proFIRE®

DNA origami traps for large viruses
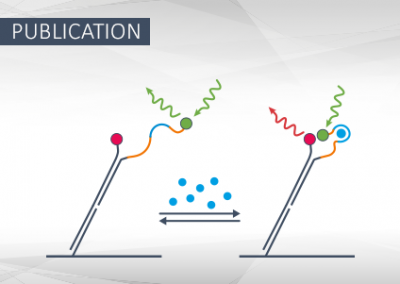
Kinetic FRET Assay to Measure Binding-Induced Conformational Changes of Nucleic Acids
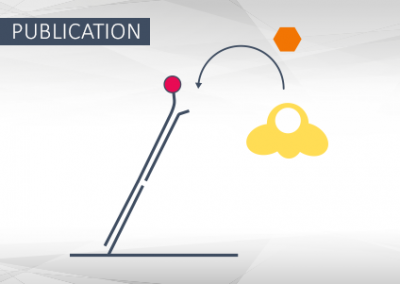
Structural and functional basis of inositol hexaphosphate stimulation of NHEJ through stabilization of Ku-XLF interaction

DISCOVER high throughput kinetic characterization of engineered multivalent peptide based inhibitors

DISCOVER protein conformational changes with electrically actuated DNA origami nanolevers

heliX Adapter Chip Storage and Handling

DISCOVER RNA protein interactions: Resolve multidomain interactions and RNA looping.

Programmable multispecific DNA-origami-based T-cell engagers
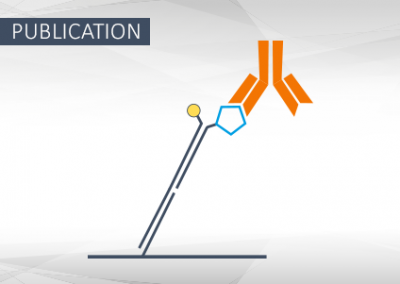
Synthetic Antigen-Conjugated DNA Systems for Antibody Detection and Characterization

DISCOVER complex membrane targets: measure binding to GPCRs, ion channels and more directly on cells